Choosing a DR Method
Since there are 16 unique methods with even more options right in this package, you may ask the question of which one you should choose. The answer can be long or short. In our paper, we detailed the various performance aspects of these DR methods, and if you need all the details, you should definitely head over there and give it a quick read. This tutorial gives you some high-level recommendations and some heuristics for you to follow.
What matters?
As our evaluation framework details, there are four major categories of accuracy metrics in DR:
Global Structure Preservation
Local Structure Preservation
Downstream Analysis Performance
scRNA-seq Concordance
While the first three are universally applicable, the last speaks to the increasing trend of integrative analysis. To read more about these evaluation criteria, head to the Evaluation Metrics section for details.
However, accuracy is not all. Since CyTOF has uniquely large sample size,
we want to ensure that each method is scalable and usable. Luckily, the
CytofDR package ensures that all the methods are user-friendly (
except for a few methods that need some TLC to be installed). In this
tutorial, we will take everything into consideration and make recommendations
accordingly.
Characteristics of Common and Top DR Methods
Here, we detail the quirks and features of our top methods! Only those that this package supports will be listed!
SAUCIE
This is a surprise: the underdog that is so likeable! But to be rational, it also has its faults!
Pro:
Great performance all around!
Mindnumbingly fast on the level of linear methods
No major faults in terms of accuracy
Con:
Software is out-of-date and its future support is question
It has some quirks and the usability suffers
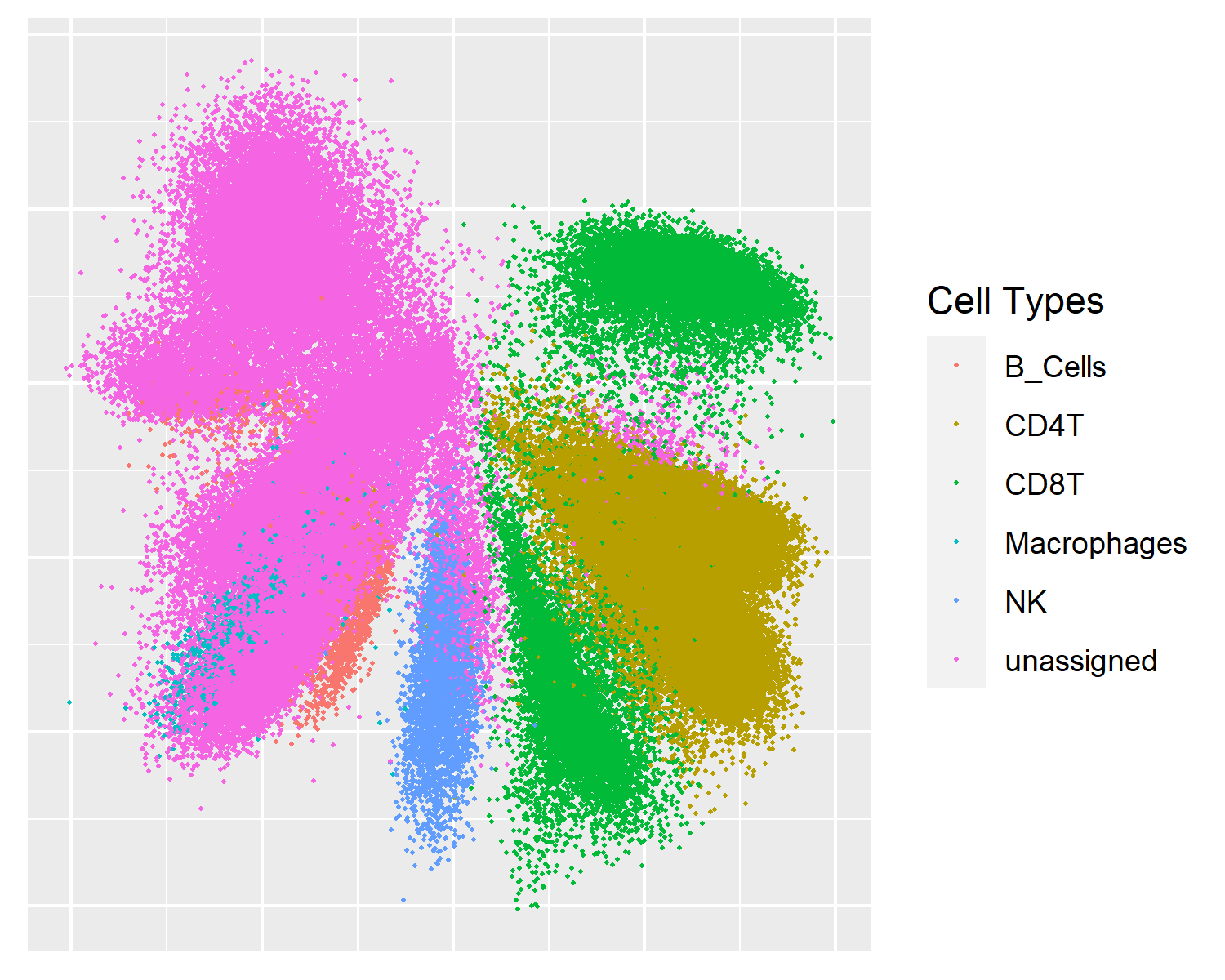
MDS
This is a complicated one! It is so traditional, and yet it presents interesting results!
Pro:
Superb global performance.
Great local performance.
Raw accuracy in terms of structural preservation is unmatched.
Con:
Dismal downstream capabilities
Almost always need downsampling for CyTOF
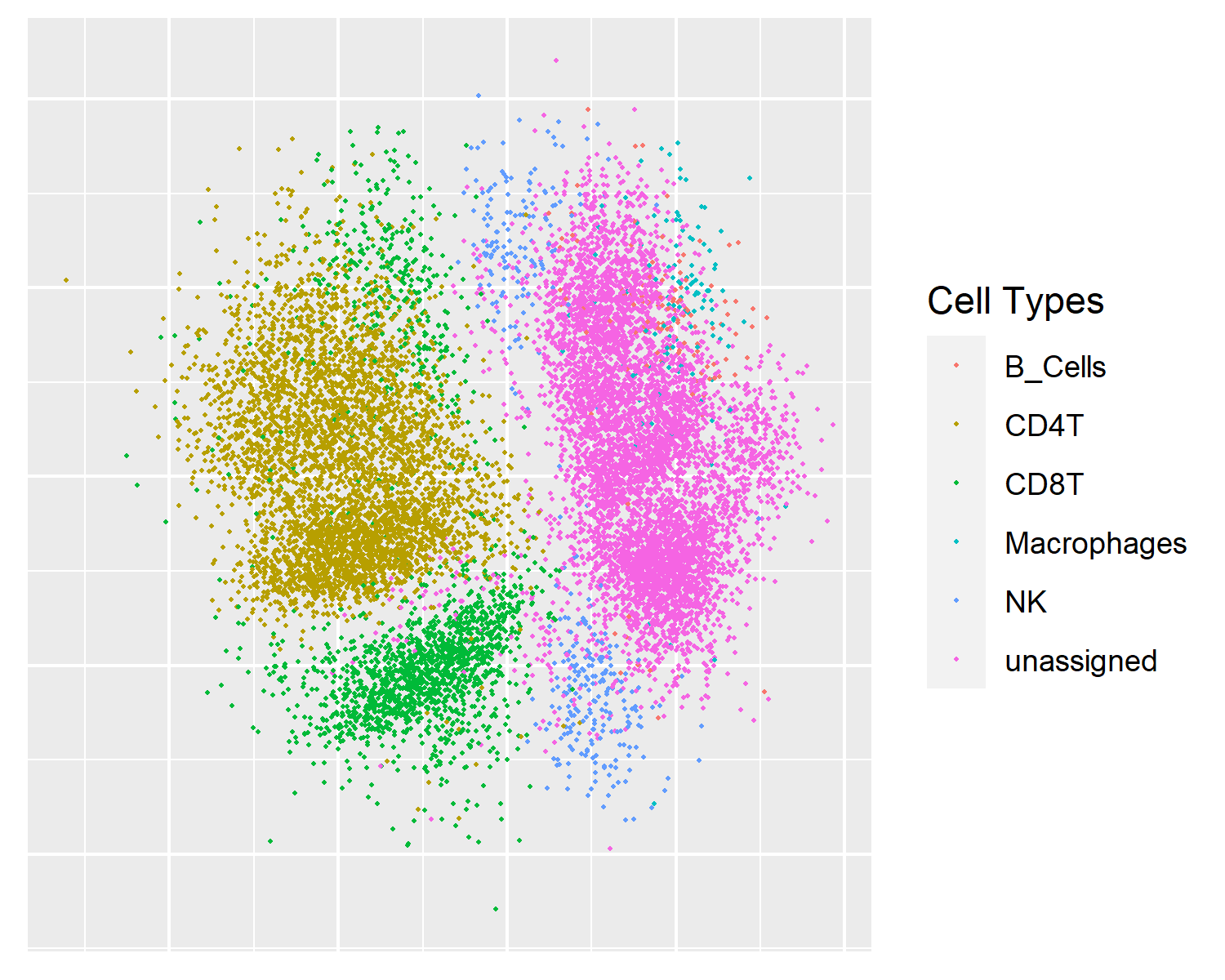
UMAP
This is the favorite among many, and it has earned its spot rightfully so.
Pro:
Superb downstream performance.
Awesome local performance.
Great documentation.
Fast and easy to use.
Con:
Global performance is not as impressive
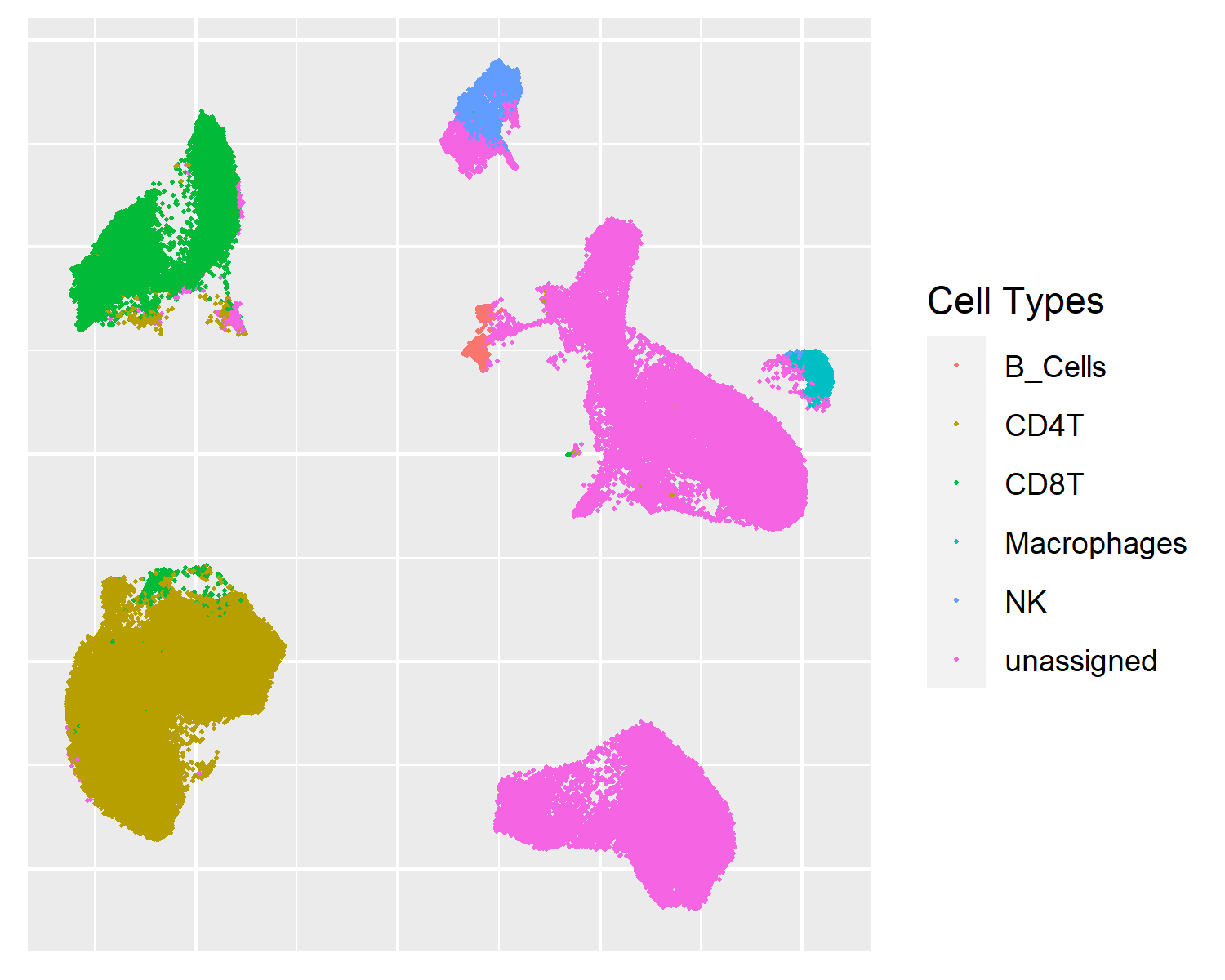
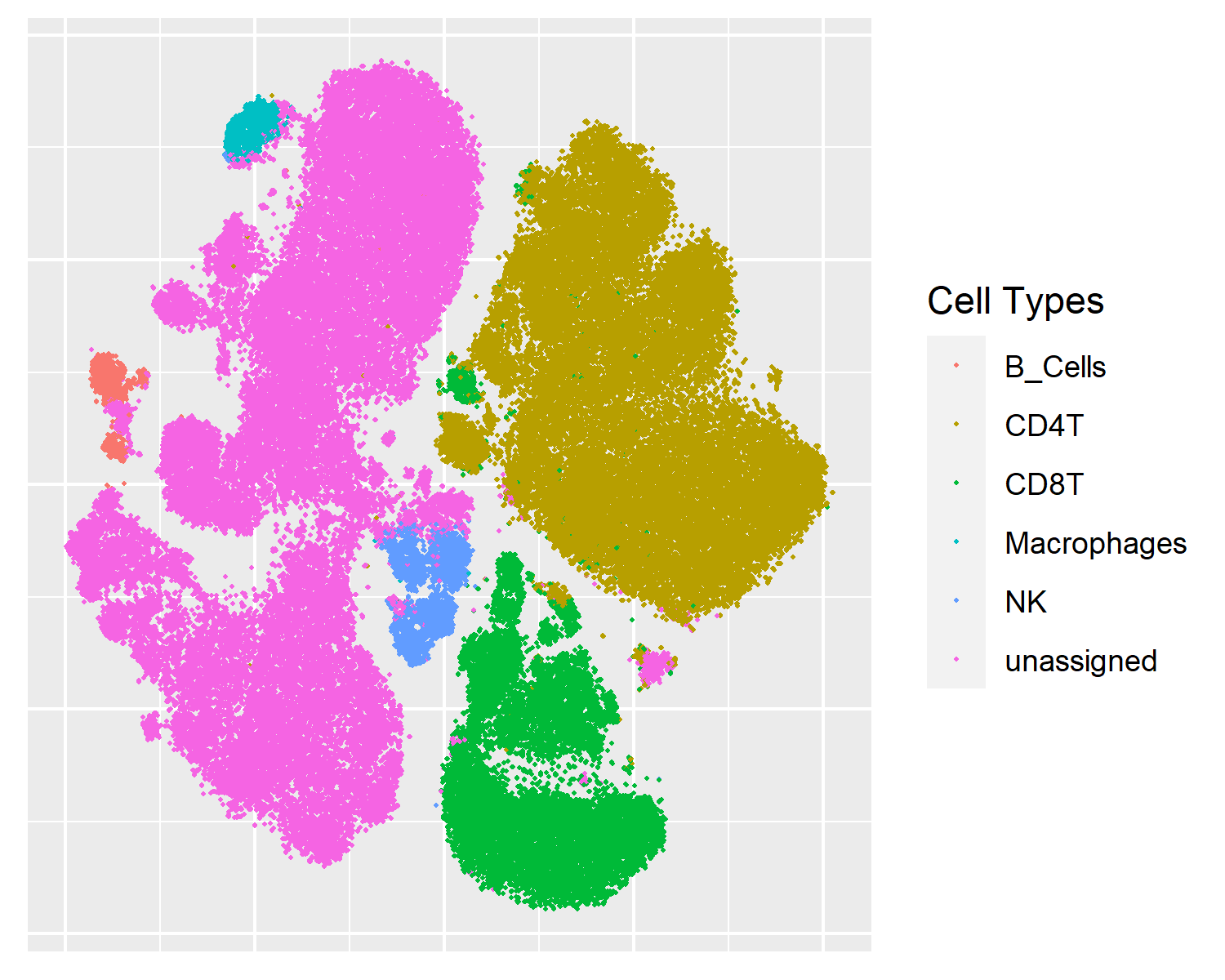
tSNE
This is the old classic: it has a loyal following despite the emergence of UMAP.
Pro:
Great local performance.
Decent downstream performance.
openTSNE is relatively fast and quite easy to use.
Con:
Global performance is rather poor
No standout feature

PCA
The linear champion! What more needs to be said?
Pro:
Great global performance.
Blazing fast
Linear and interpretable
Con:
Local and downstream are average at best
No longer the gold standard
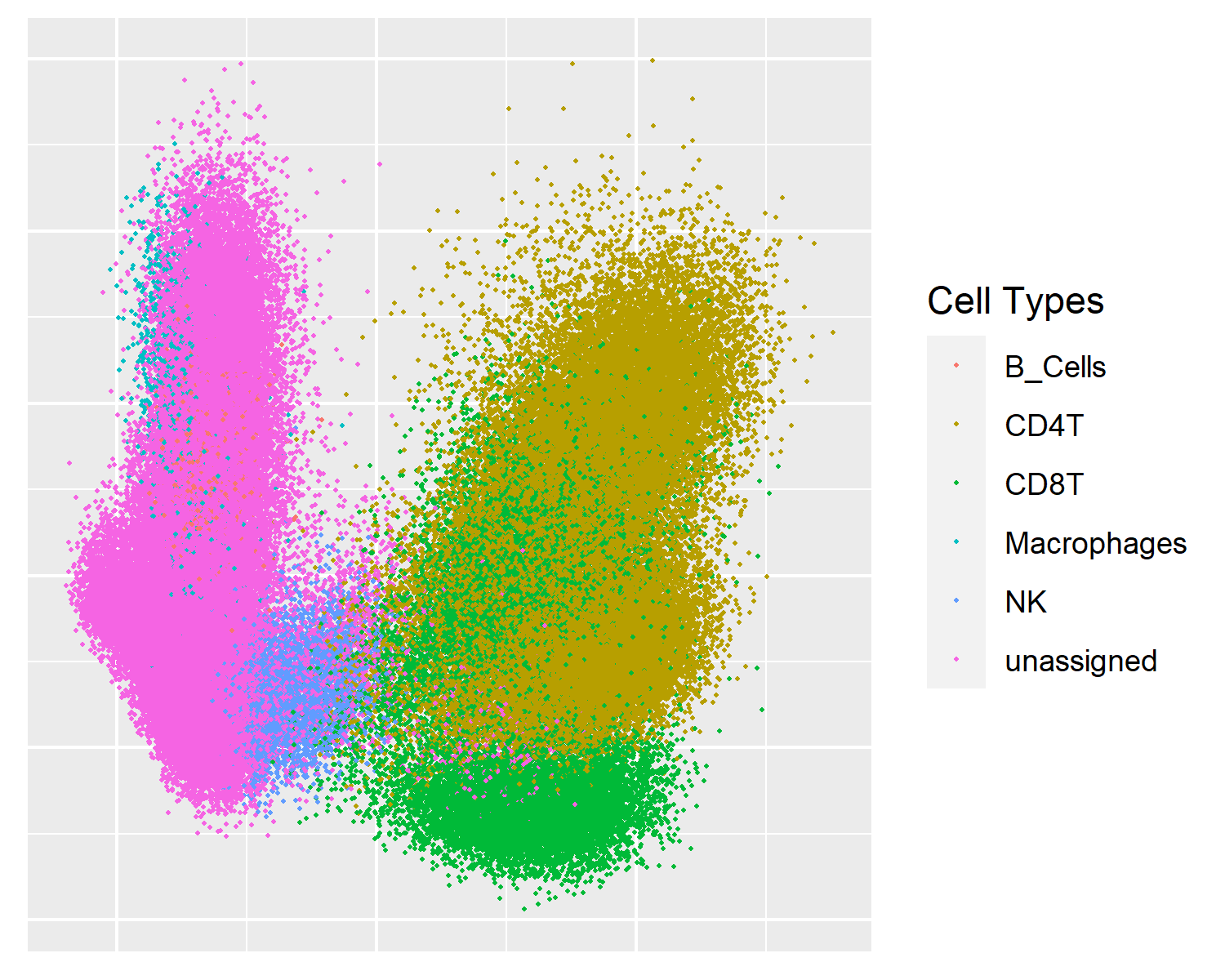
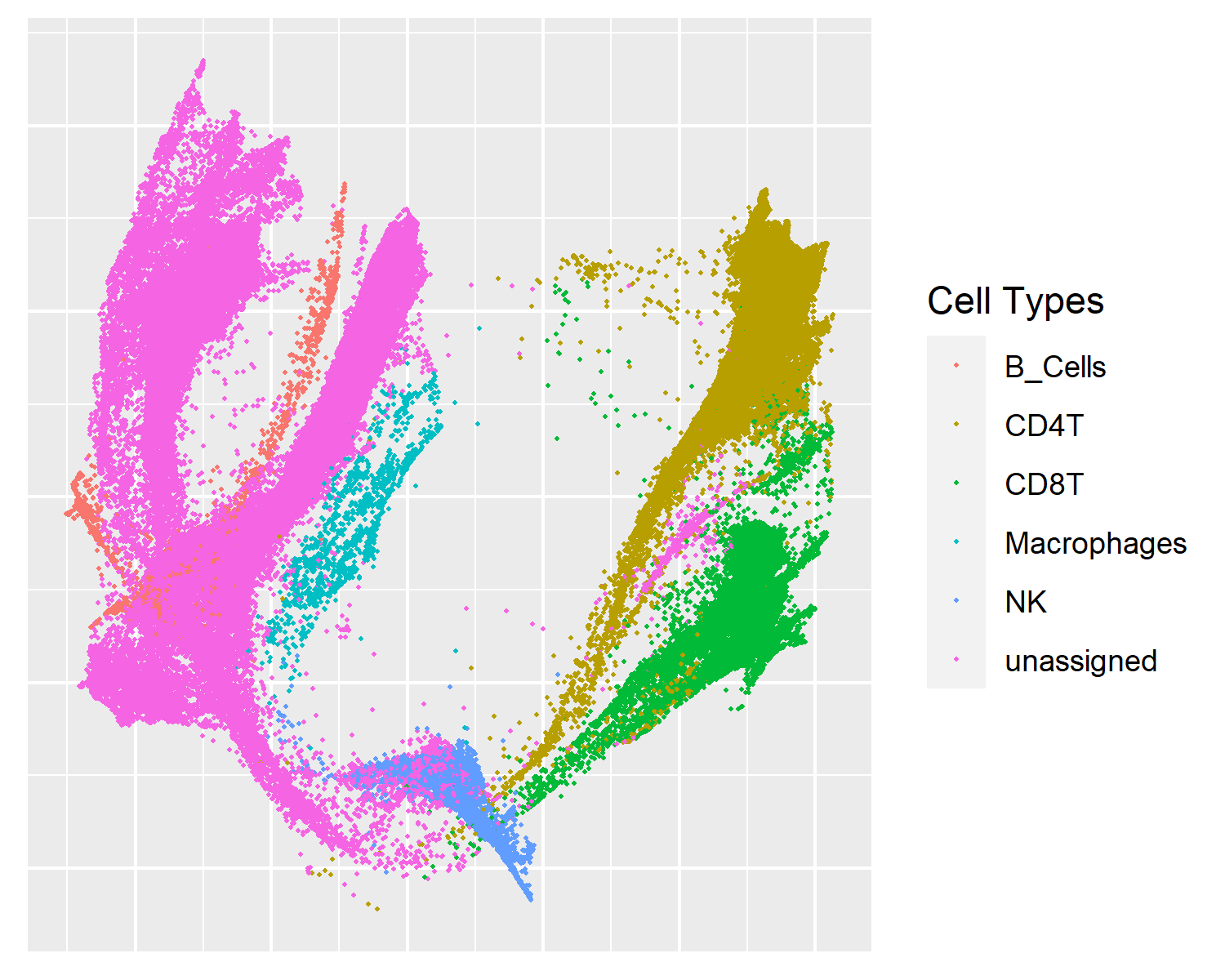
PHATE
An interesting addition to other methods with a unique focus on trajectory inference.
Pro:
Great local and downstream performance.
Embeddings uniquely resemble trajectories between clusters.
Con:
Global performance is nowhere near claimed.
Can be slow at times.

What’s your use case?
Combining all aspects of evaluation, it is very hard to come up with a
one-size-fits-all recommendation. Instead, we make recommendations
based on your use case! While CytofDR can be used in any setting,
even for other data sources that are not CyTOF, it is good to think
about what your goals are and what is important for you. In this section,
we will list a few common use cases and how you can best meet these
goals.
Further, we oftentimes recommend a couple of methods instead of just one because we strongly believe that investigating a few embeddings together will allow you to see different perspectives. As you noticed above, no one method is the best in everything. Thus, they will nicely complement each other and help you succeed.
Accuracy-Oriented Data Analysis
This is perhaps the most rigorous usecase because accuracy is of paramount importance: for example, one may want to investigate the relatonship between two cell types. In this case, we recommend that you prioritize accuracy above anything else.
In this case, you should consider MDS, SAUCIE, and UMAP. These three methods will allow you to explore both global and local performance along with good cluster resolution offered by UMAP.
Rapid Prototyping
Perhaps there is only one valid answer here: UMAP. This situation is quite common: you have a dataset and all you care is to look at an embedding with decent accuracy. UMAP is the undisputed champion to get this job done: it is cross-platform, fast, and easy to use. We would have recommended SAUCIE, if it were to be user-friendly, but its involved installation process and quirks make it unsuitable for this purpose.
Production Pipeline
This is a somewhat unique, but also important, usecase because oftentimes users want to ensure that their software pipeline is robust without worrying much about issues. In this case, we can recommend four methods here: UMAP, open_tsne, and PCA. All their packages are robust and quite well supported. Further, they’re quite scalable without needing to specifically trying to tackle the sample size issue. If you use these methods in your pipeline, you can feel at ease!
Downstream Analyses
This is closely related to the first use case, but here, we would like to highlight the clustering advantages offered by UMAP. Also, PHATE is a good option if you’re interested in investigating differentiation paths and trajectory inference. If you wish, add SAUCIE to your arsenal for its global structure preservation by compromizing a little on downstream performance, but you can always use it in conjunction with UMAP to get the best of both worlds.
I Still Can’t Choose! Help!
Well, I feel your pain! Truly, I always get too sentimental when deciding to part with certain things and choose others. In this case, just go with SAUCIE and UMAP. They are fast and will have you covered for most situations! This approach will allow you to get started and decide later if a specific need or vision arises later.